Forschung
Jorge Perez-Fernandez moved to the University of Jaén (Spain) in early 2021.
Research topic
Ribosomes are highly conserved and essential for life because they are responsible for protein synthesis. Ribosomes are formed by proteins and a special type of RNA called ribosomal RNA (rRNA). In eukaryotic cells around 78 proteins bound to 4 different rRNAs constitute the mature ribosome.
In eukaryotes, Ribosome biogenesis starts with the synthesis of the rRNA precursors by the nuclear RNA polymerases (Pols) I and III. The different pre-rRNAs are modified at several nucleotide positions and processed by several endo- and exonucleolytic cleavages to acquire the mature configuration. The incorporation of ribosomal proteins in the pre-rRNAs occurs stepwise, suggesting that these assembly events determine the final folding of the rRNAs. Most of the ribosome biogenesis steps occur in the nucleus. The pre-ribosomal particles are then actively transported to the cytoplasm where the last maturation events take place. During ribosome biogenesis more than 200 proteins interact transiently with the pre-rRNAs to guide the maturation of ribosomes. Thus, ribosome biogenesis is considered as one of the most complex processes in the cell consuming a large part of the cellular energy. In higher eukaryotes, formation of active ribosomes is highly relevant during cell growth, cell proliferation and likely differentiation. Moreover, the development of cancer cells and a novel set of diseases grouped as ribosomopathies are linked to ribosome production and function. Nevertheless, the regulation of ribosome biogenesis is far from being understood.
Our main interest is the molecular description of the early events during ribosome biogenesis. In these steps, some of the initial ribosome biogenesis factors assemble into protein subcomplexes (UTPA, UTPB, UTPC) and small nucleolar ribonucleoprotein complexes (as the U3 snoRNP). These protein complexes are coordinated with the RNA polymerase I and could drive the initial folding of the pre-rRNA. Binding of these protein subcomplexes to the pre-rRNA occurs stepwise and follows a specific hierarchy which is important for the stability of the different ribosomal precursors (fig. 1, Perez-Fernandez et al. 2007). tUTP/UTPA subcomplex is the first protein complex associating with the pre-rRNA and collaborates with RNA polymerase I to improve pre-rRNA synthesis. Binding of tUTP/UTPA is strictly required for the stable binding of UTPB and U3snoRNP. Finally, binding of UTPB and U3 subcomplexes is mutually dependent and strictly required for the subsequent binding of other ribosome biogenesis factors (Perez-Fernandez et al 2011).
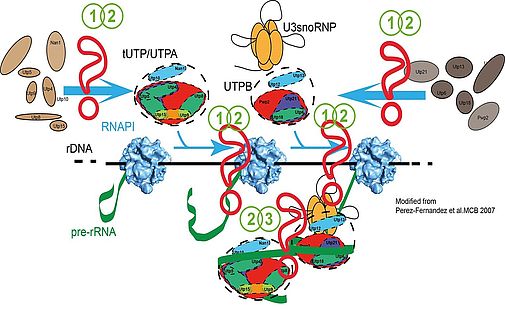
Figure 1: Principles of assembly of UTP subcomplexes. Question marks indicate pathways which are currently under investigation using different strategies which are detailed in the text. (1 and 2) in vivo formation of UTP subcomplexes and assembly into pre-ribosomal particles. (2 and 3) function during rRNA synthesis, processing and folding.
Research aims
Using the budding yeast Saccharomyces cerevisiae as model organism, we pursue a multidisciplinary strategy to contribute to a better understanding of ribosome biogenesis in eukaryotic cells.
1) Architectural studies.
Due to the high heterogeneity and the fast dynamic of initial pre-ribosomes it is difficult to analyze those structures. To solve this problem, we have developed the reconstitution of yeast UTPA and UTPB after recombinant expression of their components (Pöll et al. 2014). These initial architectural studies suggest that both subcomplexes share a common architecture based on a core complex and a dissociable dimer. Moreover, the recombinant production of UTP subcomplexes is a breakthrough which will further allow structural analysis by X-Ray and Cryo-EM.
2) In vivo assembly studies.
In vitro reconstitution of UTP subcomplexes has shown the stable association of proteins to form architectural building blocks. Moreover, our studies indicate that both complexes can be assembled independently of ongoing yeast ribosome biogenesis. The obtained structural data is used for a rational design of mutants in UTP components. The analysis of the pre-ribosomal precursors found under expression of mutant proteins constitutes the state of the art in ribosome biogenesis. For the characterization of these particles we use an approach based on semiquantitative mass spectrometry analysis (Jakob et al. 2011), conventional biochemical methods (Western blot, Northern blot...) and novel techniques developed in our department (enzymatic structure probing with MNase fusion proteins, Ohmayer et al 2011).
3) Functional studies.
Production of UTP subcomplexes is not only useful for structural analysis. A complete set of tools has been established to investigate in vitro the function of tUTP/UTPA and UTPB during formation, processing, and folding of the pre-rRNA.
Team
Jorge Perez-Fernandez | curriculum vitae |
Previous mentees | |
| Adrian Limmer, Master student 2013 | |
| Matthias Heiser, Bachelor student 2014 | |
| Fabian Hölzgen, Bachelor student 2016 | |
| Michael Brummer, Master student 2016 | |
| Christina Braun, Master student 2016, PhD student 2021 | |
| Dominik Strobel, Master student 2016 | |
| Marina Huber, Bachelor student 2017 | |
Mona Thiele, Bachelor student 2017 | |
| Tom Dielforder, Bachelor student 2018 | |
| Dietmar Sperling, Master student 2018 | |
| Catharina Schmid, Master student 2019 | |
Jorge Perez-Fernandez - Short CV
| since 2011 | Researcher at the University of Jaén (Spain) |
| 2011-2021 | Academic staff member at the University of Regensburg (Germany), Department of Biochemistry III (Chair: H. Tschochner) |
| 2007 - 2010 | Postdoctoral researcher at the Laboratory of Eukaryotic Molecular Biology (LBME, CNRS-Université Paul Sabatier, Toulouse, France) |
| 2004 - 2007 | Postdoctoral researcher at the Center for Cancer Research (CIC, CSIC-Universidad de Salamanca, Salamanca, Spain). |
| 2003 - 2004 | Postdoctoral researcher at the Center for Molecular Biology "Severo Ochoa" (CBMSO, CSIC-Universidad Autónoma de Madrid, Spain) |
| 1999 - 2003 | Doctoral Thesis (Dr. rer. nat.) at the University Autonoma of Madrid (Spain) Department of Biochemistry and Molecular Biology (Chair: JP. Garcia-Ballesta) |
Publikationen
Selected publications
Boissier F, Schmidt CM, Linnemann J, Fribourg S, Perez-Fernandez J. Pwp2 mediates UTP-B assembly via two structurally independent domains. Scientific reports. 2017, DOI: 10.1038/s41598-017-03034-y, ![]()
![]()
![]() PMID: 28600509
PMID: 28600509 ![]()
![]()
![]() %CITAVIPICKER;PMID:28600509; %CITAVIPICKER;PMID:28600509;Add to Citavi project by Pubmed ID;%Add to Citavi project by Pubmed ID;%Add to Citavi project by Pubmed ID;%
%CITAVIPICKER;PMID:28600509; %CITAVIPICKER;PMID:28600509;Add to Citavi project by Pubmed ID;%Add to Citavi project by Pubmed ID;%Add to Citavi project by Pubmed ID;% ![]()
Pérez-Fernández J, Román A, De Las Rivas J, Bustelo XR, Dosil M. The 90S preribosome is a multimodular structure that is assembled through a hierarchical mechanism. Mol Cell Biol. 2007;27: 5414–5429. doi:10.1128/MCB.00380-07 ![]()
![]()
![]()
![]()
![]()
Pérez-Fernández J, Martín-Marcos P, Dosil M. Elucidation of the assembly events required for the recruitment of Utp20, Imp4 and Bms1 onto nascent pre-ribosomes. Nucleic Acids Res. 2011;39: 8105–8121. doi:10.1093/nar/gkr508 ![]()
![]()
![]()
![]()
![]()
Jakob S, Ohmayer U, Neueder A, Hierlmeier T, Perez-Fernandez J, Hochmuth E, et al. Interrelationships between Yeast Ribosomal Protein Assembly Events and Transient Ribosome Biogenesis Factors Interactions in Early Pre-Ribosomes. PLoS ONE. 2012;7: e32552. doi:10.1371/journal.pone.0032552 ![]()
![]()
![]()
![]()
![]()
Ohmayer U, Perez-Fernandez J, Hierlmeier T, Pöll G, Williams L, Griesenbeck J, et al. Local tertiary structure probing of ribonucleoprotein particles by nuclease fusion proteins. PloS One. 2012;7: e42449. doi:10.1371/journal.pone.0042449 ![]()
![]()
![]()
![]()
![]()
Hierlmeier T, Merl J, Sauert M, Perez-Fernandez J, Schultz P, Bruckmann A, et al. Rrp5p, Noc1p and Noc2p form a protein module which is part of early large ribosomal subunit precursors in S. cerevisiae . Nucleic Acids Res. 2013;41: 1191–1210. doi:10.1093/nar/gks1056 ![]()
![]()
![]()
![]()
![]()
Ohmayer U, Gamalinda M, Sauert M, Ossowski J, Pöll G, Linnemann J, Hierlmeier, T, Perez-Fernandez, J, et al. Studies on the assembly characteristics of large subunit ribosomal proteins in S. cerevisae. PloS One. 2013;8: e68412. doi:10.1371/journal.pone.0068412 ![]()
![]()
![]()
![]()
![]()
Pöll G, Li S, Ohmayer U, Hierlmeier T, Milkereit P, Perez-Fernandez J. In Vitro Reconstitution of Yeast tUTP/UTP A and UTP B Subcomplexes Provides New Insights into Their Modular Architecture. PLoS ONE. 2014;9: e114898. doi:10.1371/journal.pone.0114898 ![]()
![]()
![]()
![]()
![]()
For full record please see the latest PubMed search results.
Kontakt
Dr.Jorge Perez-Fernandez
Departamento de Biología Experimental
Edificio B-3, 3ª Planta.
Campus Las Lagunillas s/n. 23071 - Jaén
SPAIN
T: +34-953-213036
F: +34-953-211875
E: jpfernan@ ujaen.es