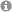Research
| Fakultät für Biologie und Vorklinische Medizin |
| Institute of Plant Sciences |
| Cell Biology and Plant Biochemistry |
| Prof. Dr. Stefanie Sprunck - Research |
In summary, our research group works on proteins contributing to cell polarity, on the gene regulatory network involved in egg cell specification, and on proteins acting on the cell surface during male-female gamete interactions.
To address questions related to our research topics we establish protocols to isolate living gametes from different flowering plant species. We perform transcriptomics and proteomics to identify candidate genes and proteins that are subsequently tested in functional studies.
Currently, we are investigating the molecular function of a small protein family that participates in the establishment and maintenance of cell polarity by characterizing specific interaction partners. Furthermore, we study RNAs and proteins putatively involved in egg cell specification, and characterize cell surface proteins which are involved in gamete adhesion, gamete activation, or fusion.
Transcriptomics of manually isolated gametes and zygotes
Transcriptomics, also referred to as expression profiling, is the investigation of a set (or all) RNA molecules expressed in a certain tissue or cell type, in a specific developmental stage, or under certain physiological conditions. Besides expression profiling, transcriptomics serves as a screening tool for candidate gene discovery.
We are manually isolating living cells from flowering plant female and male gametophytes by micromanipulation with the aim to study the transcriptome of these cells. Previously, we used female gametes (egg cells and central cells) and two-celled proembryos of wheat and Arabidopsis to perform either single-run partial sequencing of randomly selected cDNA clones (ESTs; “expressed sequence tags”) of generated cDNA libraries (Sprunck et al., 2005), or microarray-based expression profiling (Englhart et al., 2017). Meanwhile, we perform next-generation sequencing (NGS) to investigate the RNA and small RNA content of female and male gametophytic cells from different flowering plant species.
We are analyzing our transcriptome data sets with the aim to discover uncharacterized gamete-specific or early fertilization-induced mRNAs, as well as small and long noncoding RNAs. We identified, for example, transcripts specifically present in the egg cell and/or the central cell and transcripts specifically induced in zygotes and two-celled proembryos (Chen et al., 2017; Leljac-Levanic et al., 2013; Sprunck et al., 2012; Juranic et al., 2012; Sprunck et al., 2005). Currently, we investigate the molecular function of selected candidate genes/gene families participating in the establishment of cell polarity, in gamete specification, and in gamete interactions during double fertilization.
Establishment of cell polarity
Establishing and maintaining proper cell polarity are key aspects of life. One of the major goals of cell biology is to understand the molecular mechanisms mediating cell polarization. Generally, cell polarity means cellular asymmetry.
Polarization and asymmetrical growth in eukaryotic cells require elaborate regulation of a variety of cellular processes, including spatially restricted signaling, reorganization of the cytoskeleton, and polarized membrane trafficking.
Polarized cells divide asymmetrically, generating daughter cells that are different in cellular contents, cell size, or developmental potential.
The asymmetric distribution of cell fate determinants (e.g., proteins and RNAs) but also asymmetric translational repression and asymmetric protein degradation govern cell fate determination and differentiation, which is essential for the development of a multicellular organism or tissue, containing cells of different molecular fate.
To study the establishment and maintenance of cell polarity in flowering plants we use the following Arabidopsis model cells: pollen tubes, root hairs, trichomes, egg cells/zygotes.
1. Polar tip growth of pollen tubes
The pollen tube is among the fastest growing plant cells. It is widely used as a model cell to study tip growth.
The pollen grain germinates on the surface of the stigma and forms an elongating tube that navigates through the female tissues toward the ovule to deliver its two sperm cells for double fertilization. Like growth cones of axons, pollen tubes require a signaling network to recognize and interpret extracellular cues that determine the site and the direction of tip growth.
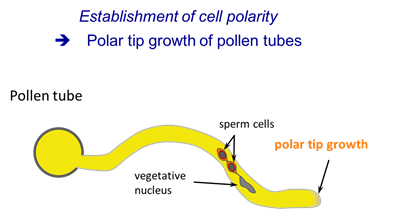
Within the past years, crucial molecular players of the pollen tube tip growth machinery have been identified, including a tip-focused calcium gradient, elaborate F-actin dynamics, tip-directed vesicle trafficking, and localized exocytosis. In our lab, we identified the Armadillo repeat protein ARMADILLO REPEAT ONLY 1 (ARO1) as a novel player of the pollen tube tip growth machinery (Gebert et al., 2008). Currently, we investigate the mechanistic function of ARO1 and related ARO protein family members in cell polarization.
2. Establishment of polarity in the egg cell
The egg cell and the zygote
The mature egg cell of the model plant Arabidopsis thaliana is a highly-polarized cell. The large vacuole is located at the micropylar pole of the egg cell, adjacent to the micropyle of the ovule (the entrance point of the pollen tube), while the nucleus and most of the cytoplasm is located at the opposing chalazal pole.
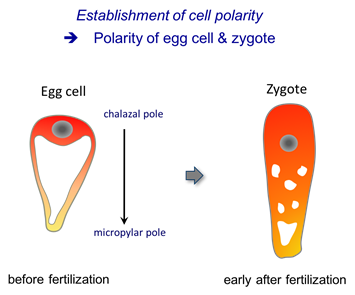
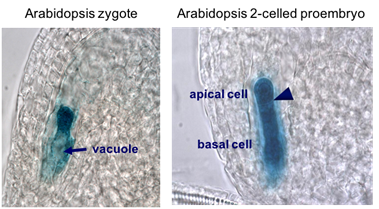
Immediately after fertilization, the vacuolar organization changes and the zygote nucleus is moved away from the chalazal pole. Live-cell imaging during the entire process of zygote polarization revealed that the preexisting alignment of microtubules and F-actin in the egg cell is lost on fertilization and that the rapidly elongating zygote reorients the cytoskeletons to perform directional cell elongation and polar nuclear migration (Kimata et al. 2016, PNAS 113, 14157-14162).
The first zygotic cell division is transverse and asymmetric, producing a smaller apical cell and a larger basal cell. The two daughter cells do not only differ in terms of their morphology but also in their developmental fate. The apical cell gives rise to the later embryo, while the basal cell mainly forms the extra-embryonic suspensor.
3. ARO proteins and polarity
In order to identify novel players participating in the establishment of cell polarity in egg cells and pollen tubes, we performed transcriptomics approaches using isolated egg cells of wheat and Arabidopsis. We focused on uncharacterized (novel) transcripts present both in egg cell and in pollen tubes.
One of the first candidates chosen for functional analyses was a transcript encoding the Armadillo repeat protein ARMADILLO REPEAT ONLY 1 (TaARO1) and its putative ortholog in Arabidopsis (AtARO1).
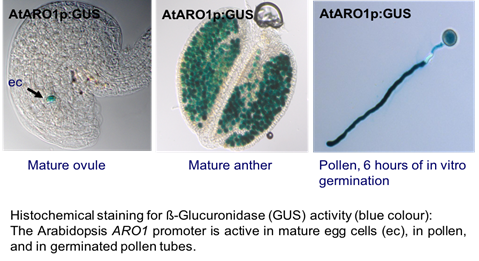
ARO1 genes are specifically expressed in the pollen vegetative cell and the egg cell. In the Arabidopsis pollen tube, the AtARO1-GFP (green fluorescent protein) fusion protein localizes to vesicles, accumulating in the inverted cone-shaped region of the pollen tube tip (Vogler et al., 2015). Moreover, the accumulation of AtARO1-GFP in the pollen tube tip is sensitive to brefeldin A (BFA), a fungal metabolite that inhibits protein secretion. These observations support our hypothesis that the observed partial co-localization of AtARO1-GFP with F-actin in the shank of pollen tubes reflects the vesicle transport along longitudinal F-actin cables (Gebert et al., 2008).
We found AtARO1 to be of fundamental importance for polar pollen tube tip growth. Compared with wild type pollen tubes, the pollen tubes of AtARO1 knockout plants (aro1-3) revealed a highly-disorganized F-actin cytoskeleton and growth depolarization (Gebert et al., 2008).
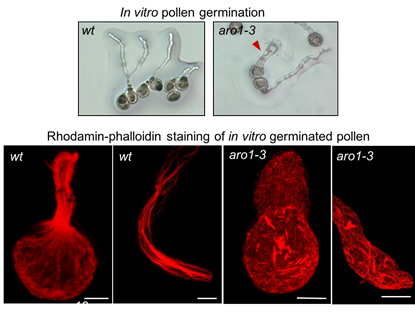
No phenotype was observed in egg cells, suggesting functional redundancy within the ARO family members. The role of ARO proteins for egg cell and/or zygote polarity is currently being analyzed in mutants where we knocked out the whole gene family.
The main aims and research questions of this project are: - How do the ARO proteins act on the molecular level? What other cellular components and proteins interact with ARO proteins? - Do we find similar phenotypes and interaction partners in egg cells/zygotes and pollen tubes, indicating that the molecular mechanisms of cell polarity formation in both cells follow the same paradigm?
Gamete interactions during double fertilization
1. Female gametophyte development and double fertilization
Plant gametes are, in contrast with animal gametes, not direct products of meiosis but differentiate within multicellular haploid generations, the male and the female gametophytes. In flowering plants (angiosperms), the gametophytic generations are reduced to a few cells that are located within the diploid sporophytic tissues of the flower.
The female gametophyte (or embryo sac) is enclosed by the ovule that is located within the ovary. In Arabidopsis, the female gametophyte originates from a single haploid spore (functional megaspore), which undergoes three free nuclear divisions to develop into an eight-nucleate syncytium. Following cellularization, the distinct haploid cell types of the female gametophyte are formed: Two female gametic cells, the egg cell and the central cell, which are flanked by accessory cells. The large vacuole of the central cell occupies the central region of the female gametophyte. Two synergid cells neighbour the egg cell at the micropylar end of the female gametophyte, while three antipodals are adjacent to the central cell at the chalazal end (Sprunck & Groß-Hardt, 2011).
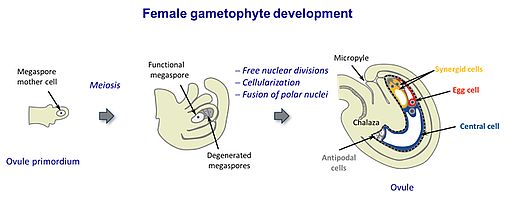
Double fertilization
Fertilization in flowering plants is unique among all known organisms in that not one, but two, female reproductive cells are fertilized in a process called double fertilization. A complex mechanism involving two male gametes (sperm cells) and two female gametes (egg cell and central cell) results in two distinct fertilization products, the diploid embryo and the triploid endosperm. Both fertilization products are required to achieve successful seed development.
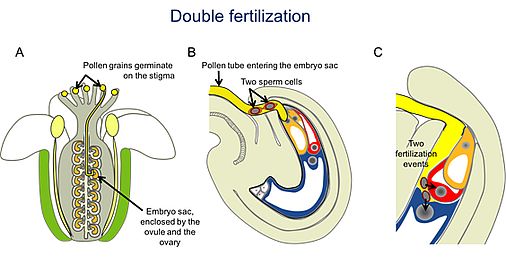
During double fertilization, the two non-motile sperm cells are transported as a pair by the growing pollen tube, which is guided through the style and the transmitting tract of the pistil to the ovule (A). Finally, the pollen tube grows through the micropyle of the ovule into the embryo sac (B), where two female gametes (egg cell and central cell) are located. After pollen tube burst, the sperm cell pair is released towards a cleft between the egg and central cell. One sperm cell fuses with the egg cell, while the second sperm cell fuses with the central cell (C), resulting in two distinct fertilization products: the embryo and the endosperm.
In contrast with the molecular events involved in guided pollen tube growth and sperm cell delivery into the female gametophyte, almost nothing is known about later processes, covering the phase from sperm cell release until gamete fusion (plasmogamy) (Sprunck, 2010). Within minutes after a sperm cell pair is delivered, each one of the sperm cells fuse with one of the female gametes to initiate seed development in a coordinated manner. Attempts to elucidate these issues at the cellular and molecular levels are hampered by the fact that angiosperm gamete interactions are short-lived events, taking place deeply embedded in the maternal tissues of the ovule and ovary.
2. Monitoring male-female gamete interactions in Arabidopsis
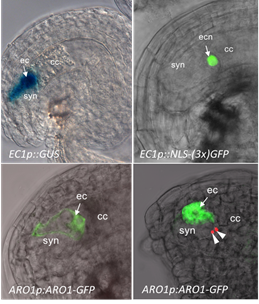
Using our expression data derived from egg cells and central cells from wheat (Sprunck et al., 2005), we identified gamete-specific genes and their putative orthologs in Arabidopsis. We used the corresponding promoter regions to generate ß-glucuronidase (GUS) and green fluorescent protein (GFP) marker lines for the Arabidopsis egg cell, the central cell, and synergid cells. The green fluorescent ARO1 marker (Gebert et al., 2008) labels the egg cell cytoplasm (EC) in ovules at maturity, while the green fluorescent EC1.1p:NLS-3xGFP marker is specifically expressed in the egg cell nucleus (ECN). These and new membrane marker lines now serve as important tools to investigate gamete interactions on the cellular and molecular level. The interaction of the gametes can be observed microscopically, when our marker lines are pollinated with sperm cell membrane marker lines (Sprunck et al., 2012), or the previously published red fluorescent sperm nucleus marker line HTR10-mRFP1 (Ingouff et al., 2009).
Currently, these marker lines are used to study mutants that have been generated in our group, showing defects during double fertilization.
3. Cell surface molecules involved in gamete recognition and/or fusion
Male-female gamete recognition, adhesion, and fusion encompass the most intimate steps during fertilization. Nevertheless, the knowledge about molecules that act on the cell surface of flowering plant gametes is scarce.
We identified a cluster of egg cell-specific transcripts in our expression data from wheat egg cells (EC-1) that encode secreted small cysteine-rich EC1 proteins (Sprunck et al., 2005). Five genetically redundant EC1 genes (EC1.1 to EC1.5) are present in the genome of Arabidopsis thaliana. Functional studies revealed that egg cell-secreted EC1 proteins are essential for the sperm cells to gain fusion competence (Sprunck et al., 2012, Rademacher and Sprunck, 2013).
Currently, we are analyzing our transcriptome and proteome data to identify more cell surface proteins which are involved in male-female gamete recognition, adhesion, and fusion.
The main aims and research questions of this project are:
- Which molecules on the cell surface of female and male gametes mediate gamete recognition, adhesion and fusion in flowering plants?
- How do the egg cell-secreted EC1 proteins act on the molecular level?
- Are there species-specific gamete interaction mechanisms in flowering plants?
Methods
We are using a wide range of basic and advanced methods related to (plant) molecular biology, molecular genetics, cell biology and protein biochemistry.
- DNA work (DNA isolation, conventional & Gateway®-based cloning, PCR, real-time quantitative PCR, PCR-mediated mutagenesis, Overlap extension PCR cloning, RNAi, amiRNA, CRISPR/Cas9, Southern Blot, Southwestern Blot, cDNA libraries, etc.)
- RNA work (total RNA and mRNA isolation, small RNA isolation, cDNA synthesis, RT-PCR, In vitro-transcription, DIG labeling, In situ hybridization, Northern Blot, etc.)
- Protein work (protein extraction, PAGE, 2D-PAGE, Western Blot, protein staining, immunoprecipitation, protein phosphorylation, recombinant protein expression, affinity purification, Size Exclusion Chromatography, Äkta pure system, etc.)
- Interaction studies (Y1H and Y2H screens, EMSA, Co-immunoprecipitation, etc.)
- Microscopy (fluorescence microscopy, DIC microscopy, Confocal Laser Scanning Microscopy, Spinning-disc live-cell imaging, immunofluorescence, etc.)
- Transformation of bacteria and yeast
- Plant transformation (biolistic, Agrobacterium-mediated) and genetics
- Plant cell culture (Arabidopsis, wheat, maize)
- Micromanipulation (single cell isolation)
- Expression studies (Microarrays, RNAseq, RT-PCR)
Prof. Dr. Stefanie Sprunck
Cell Biology and Plant Biochemistry
University of Regensburg
Universitätsstraße 31
93053 Regensburg
Germany
Phone: +49 (0)941 943-3005
Fax: +49 (0)941 943-3352
E-Mail