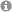Research
Research at our department
Sequencing of the complete human genome revealed that only a very small proportion of the genetic information is protein coding. Approximately 90% of the human genome, however, is not protein coding and was initially considered to be some kind of evolutionary ‘junk-DNA’. Today, we know that almost all non-coding DNA is actively transcribed to RNA and that such RNA molecules are very important for many cellular processes. Intensive research during the last years revealed that non-coding RNAs are involved in the regulation of different diseases like neurodegenerative diseases or cancer. The investigation of these RNAs is just at its beginning and future studies will certainly result in new prospects for cancer research.
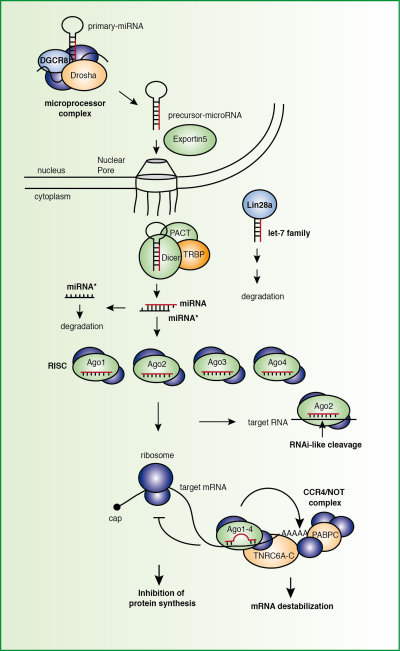
Our research is focused on the investigation of non-coding RNA pathways in mammalian cells. Small RNAs such as short interfering RNAs (siRNAs) or microRNAs (miRNAs) are processed from double stranded (ds) precursor molecules by the action of RNase III enzymes. MiRNAs utilize the enzymes Drosha and Dicer while siRNAs typically only need Dicer processing from long dsRNA species. Both classes are single stranded small RNAs that are incorporated into the RNA-induced silencing complex (RISC). The direct binding partner of the small RNA is a member of the Argonaute protein family, which has been subject of our research for the past ten years. In current projects, we investigate posttranslational modifications of key miRNA biogenesis as well as effector proteins. Which signaling pathways and which kinases or phosphatases act on these factors? What are the biological consequences of phosphorylation at specific sites? These are two key questions in this project (see 1).
In parallel to the biochemical characterization of small RNA pathways in mammalian cells, we use RNA cloning and deep sequencing to profile short and long non-coding RNAs in cancer. In close collaboration with the medical faculty (Christina Hackl), we investigate non-coding RNAs in colorectal cancer progression. Together with Peter Hau, we profile non-coding RNA in glioblastoma patients (see 2).
In addition, we have started to identify and characterize RNA binding proteins that interact with specific miRNA precursor molecules. Using biochemical approaches, we identified hundreds of such proteins that sequence-specifically bind to pre-miRNAs. We are currently investigating the biological functions of these proteins. We employ state-of-the-art technologies such as CRISPR/Cas9 knock out or in vitro selection of RNA binding motifs (bind’n’seq). We also started to investigate novel RNA binding proteins and RNA binding domains (see 3).
Finally, we characterize specific modifications on RNA molecules. The availability of antibodies against m6Adenin (m6A) allowed for a better characterization of this modification. Interestingly, this modification is highly abundant on mRNAs and many interesting biological functions of this modification have been reported. In our projects, we characterize enzymes and RNA binding proteins that act in this modification pathway. In addition, we generate monoclonal antibodies against a set of ten different base modifications. With these tools, we hope to map such modifications on RNA molecules and generate a dynamic view on the biology of RNA base modifications (see 4).
1 Post-translational regulation of small RNA pathways
Post-translational modifications are found on many proteins and contribute to functional diversifications of gene products. It is the aim of our work to catalogue and functionally characterize post-translational modifications on the key-components of the miRNA pathway. We use state-of-the-art quantitative mass spectrometry to identify and absolutely quantify modifications such as methylation or phosphorylation. For biologically relevant investigations, we have generated monoclonal antibodies against all key components of the miRNA pathway. We map phosphorylation sites on factors of the miRNA pathway (Drosha, Xpo-5, Lin28) and functionally characterize the identified phosphorylation sites on Ago and TNRC6 proteins. Signaling pathways and phosphorylation-dependent interaction partners will be investigated.
|
| |
| Figure 1: The mammalian miRNA pathway. Post-translational modifications on key components of the pathway will be mapped and functionally characterized. |
2 Profiling of non-coding RNAs in cancer
MicroRNAs (miRNAs) are important gene regulators. They interact with distinct binding sites on mRNAs leading to inhibition of gene expression. Depending on the target they regulate, miRNAs can function as oncogenes or tumor suppressors. Long non-coding RNAs (lncRNAs) have been implicated in various diseases including cancer. A model of a novel ‘RNA language’ based on competing endogenous RNAs (ceRNAs) has been put forward. ceRNAs compete with mRNAs for miRNAs and can regulate whole transcriptomes. However, nothing is known about the interplay between miRNAs and lncRNAs during metastasis and therapy. Very recently, circular RNAs have been found and we are characterizing several interesting circular RNA molecules during cancer progression.
In this project, we use different mouse models (BALBneuT and an orthotopic model of colorectal cancer) and analyze miRNA, lncRNA and circular RNA expression during cancer progression and therapeutic intervention by deep sequencing. Changes of non-coding RNA expression between the primary tumor and metastases will be characterized. MiRNA targets including putative ceRNAs will be identified and their relevance will be analyzed in vitro and in vivo.
3 Regulation of gene expression by RNA-binding proteins
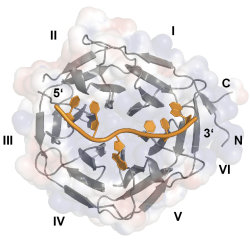
EEACoding and non-coding RNAs are regulated at various different levels. For example, microRNA (miRNA) expression can be influenced at multiple steps of biosynthesis. Furthermore, it has been reported that direct RNA methylation events can also affect gene expression in many different organisms and systems. In some of our projects, we aim at identifying and characterizing factors that affect the maturation of miRNAs. We employ biochemical pulldown assays to isolate specific binding partners of different miRNA species. We analyze the physiological role of these factors using molecular or cell biological approaches as well as the molecular details of miRNA-protein interactions using x-ray crystallography and other biochemical and biophysical methods. One example of our approach is highlighted by our recent characterization of the RNA binding activity of Drosophila Brain tumor (BRAT). This protein has not been implicated in RNA binding before and we showed by x-ray crystallography, that the NHL domain is a novel RNA binding domain. We hope to identify additional novel RNA binding domains in the future.
|
| |
| Figure 2: Structure of the NHL domain of BRAT in complex with its RNA binding motif (Loedige et al. 2015). |
4 Investigation of RNA-specific base modification pathways
RNA base modifications have been known for a long time but only recently it was recognized that such modifications are also found on mRNAs and the type and extent of modification can have dramatic effects on gene expression. The underlying mechanisms, however, are largely unknown. One specific type of such modifications is m6A methylation. We decipher the role of the m6A methylation pathway on the regulation of coding and non-coding RNAs. Writers, readers and erasers of this modification have already been identified. However, not much is known about the composition of specific protein complexes as well as the atomic structure of these factors. Thus, it is the aim of our projects to functionally and structurally characterize known and putative novel factors of the m6A methylation pathway, which we aim to identify in human cells.
Recently, additional base modifications have been identified on mRNA molecules. These include 5-methylcytidin or pseudouridin and it has been shown that such modifications influence the functions of target RNAs as well. Likewise, many more of chemically known RNA modifications may reside on mRNAs but due to the lack of reagents have not been identified so far. Thus, we plan to generate such tools in the first part of our proposal. We will establish monoclonal antibodies against a variety of known base modifications and will use them for characterizing RNAs carrying such modifications. In the second part of the proposal, we plan to investigate so far uncharacterized putative RNA methyltransferases of the METTL protein family. Using homology search and structure modeling, we found that several members can be sub-grouped (e.g. METTL2A/B, 6 und 8) and it is likely that these proteins are functionally related and may have similar RNA targets. We will use biochemical approaches to identify potential interaction partners (RNA and protein). Finally, we will also investigate these protein or protein domains structurally using x-ray crystallography. With this research proposal, we will help to better understand the complex regulation that is mediated by RNA modification.