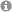Research
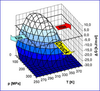
The functions of proteins rely on their structures and the structural changes they can undergo. Structural biology hence provides unique insights into biomolecular function and thereby into cellular processes. Furthermore, the knowledge of protein structures is an invaluable starting point in the development of drugs to treat human disease. Nuclear magnetic resonance (NMR) spectroscopy is the only method available to study biomolecular structure and dynamics with atomic resolution in solution. NMR provides information on the three-dimensional fold of proteins and the way proteins interact with other biomolecules. Finally, NMR spectroscopy gives insights into how protein complexes move and change conformation, aspects that are important for many enzymatic reactions. The information gained by NMR about biomolecular structure, interactions and dynamics allows us to understand how proteins are able to perform specific tasks in the cell.

left to right: Silke Wiesner, Nadine Stephan, Jobst Liebau, Philip Wurm, Julian Huebner, Johannes Schmoll, Daniela Lazzaretti, Patricia Albrecht, Simon-Peter Grunert, Alex Schmalix, Zeliha Koca, Christina Krempl, Kinga Ay, David Stelzig, Remco Sprangers.
TEAM
Phone | room | Email | ||
Team Leaders | ||||
| Prof. Dr. Remco Sprangers | 7751 | 7.2.07 | ||
| Prof. Dr. Werner Kremer | 2185 | 7.2.09a | ||
| Dr. Silke Wiesner | 7752 | 7.2.25 | ||
| Prof. Dr. Dr. Hans-Robert Kalbitzer (emeritus) | 2594 | 7.1.22 | ||
SECRETARIAT | ||||
| Patricia Albrecht | 2194 | 7.2.08 | ||
Technical Assistants | ||||
| Nadine Stefan (wet lab) | 7753 | 7.2.06 | ||
| Simon Peter Grunert (NMR) | 4181 | 7.2.24 | ||
Post-docs | ||||
| Jan Overbeck | 2491 | 7.2.06 | ||
| Daniela Lazzaretti | 2491 | 7.2.06 | ||
| Jobst Liebau | 2495 | 7.2.05 | ||
| Julian Hübner | 2495 | 7.2.05 | ||
| Christina Krempl | 2491 | 7.2.06 | ||
PHD Students | ||||
| Alexander Schmalix | 4181 | 7.2.24 | ||
| David Stelzig | 4181 | 7.2.24 | ||
|
| ||||
Masters students | ||||
| Zeliha Koca | 7754 | 7.2.24 | ||
Bachelor students | ||||
| Thomas Wolters | 7754 | 7.2.24 | ||
visiting students |
contact
Address:
University of Regensburg
Universitätsstrasse 31
93053 Regensburg
How to get to us?
Due to ongoing construction sites on the campus, you might like to consult the university's navigation system UR walking.
If you prefer a map, look here.
Our labs and offices are located on the second floor.
Postal Address:
University of Regensburg
93040 Regensburg
Secretary:
Room 7.2.08
Phone +49-941-943 2194
Fax +49-941-943 2479
Overview of Telephone Numbers:
Office Remco Sprangers (7.2.07): 7751
Office Silke Wiesner (7.2.25): 7752
Office Werner Kremer (7.2.09a): 2185
Sekretariat (7.2.08): 2194
Office Team Sprangers I (7.2.06): 2491
Office Team Sprangers II (7.2.05): 2495
Office Team Wiesner (7.2.24): 4181
Lab Sprangers (7.2.03/04): 7753
Lab Wiesner (7.2.22/23): 7754
500 MHz Spectrometer: 2184 (only from inside University)
600 MHz Spectrometer: 4183 (only from inside University)
800 MHz Spectrometer: 2848 (only from inside University)
NMR Operating And Usage Concept
publications
Vacancies
Bachelor and Masters theses
Students interested in doing their Bachelor or Masters thesis in the Sprangers or Wiesner lab are welcome to contact us. There is a range of possible projects with a combination of structural biology and biochemical and biophysical methods.
PhD students and Postdocs
We have an open position for a Postdoc and for a PhD student.
Our lab is interested in applying and developing methods in NMR spectroscopy (including methyl TROSY techniques) for very large protein complexes (> 100 kDa). Our aim is to understand how motions in large enzyme complexes influence function.
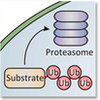
Methodologically, we combine high resolution NMR spectroscopy with other structural techniques (X-ray crystallography, cryo-EM), biophysical methods (including ITC, fluorescence anisotropy), activity measurements and protein/RNA chemistry. From a biological point of view our interests are enzymes that are involved in the degradation of mRNA.
Our wetlabs and offices are newly renovated and fully equipped for modern molecular biology methods. In addition, we run three NMR spectrometers (800, 600 and 500 MHz) are equipped with new consoles and cryo/prodigy prodeheads.
Regensburg is a lively and attractive UNESCO world heritage city with a high density of pubs and bars. The faculty of biology at the Regensburg university has a focus on RNA biology.
We are looking for enthousisatic PhD/ Postdoc candidates with a strong interest and background in biomolecular NMR spectroscopy. In addition, experience in standard molecular biology techniques (cloning, protein expression and purification) is a strong plus.
Interested candidates should contact me directly (remco.sprangers@ur.de) and include names of two peers that can provide letters of recommendations. In addition, a concise description of skills, previous work experiences and scientific interest should be included.