SFB960 - Research
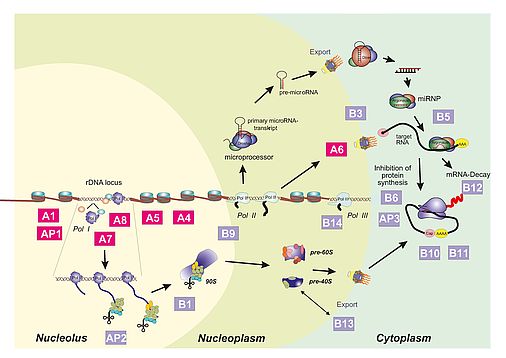
Part A: (r)RNA synthesizing machineries and chromatin
Project A1: H. Tschochner, J. Griesenbeck, P. Milkereit
Analyses of the RNA polymerase I (Pol I) machinery: specific transcriptional mechanisms, comparison with other RNA polymerases and structural determination of Pol I complexes
RNA polymerase I (Pol I) is a specialized enzyme which transcribes ribosomal RNA (rRNA) genes in all eukaryotes investigated so far. Pol I transcription is distinguished by specific DNA cis-elements and trans-acting factors as well as the usage of a specific chromatin template. The aim of this project is to unravel specific functional and structural features of the Pol I machinery on a molecular level and to get more insights into the mechanism of Pol I dependent RNA synthesis. Well-defined in vitro systems, which were established in the first funding period will now be used to investigate how certain subunits, domains and/or posttranslational modifications of the Pol I machinery are involved in specific steps of the transcription cycle.
Click here for more information.
Project A4/B2: G. Längst
Coordination of rRNA gene transcription initiation and termination in chromatin
Non-coding RNAs are regulatory molecules in the cell that in part are exerting their function bound to chromatin. We showed that the formation of triple helices is one mechanism of sequence specific targeting of RNA to chromatin and that these structures exist in vivo. We will first, address the molecular function of triple helix bound RNAs on gene regulation and chromatin structure, and second analyse the function and structure of our newly characterized triple helix binding proteins involved in rRNA gene regulation.
Click here for more information.
Project A5: J. Griesenbeck, P. Milkereit, H. Tschochner
Analysis of ribosomal RNA gene chromatin in distinct transcriptional states
The essential process of ribosomal RNA (rRNA) gene transcription in the nucleolus of eukaryotic cells occurs in the context of chromatin. Our project aims to define rRNA gene chromatin, in its composition, structure and function. In the last funding periods we have further developed a technique for the native isolation of rRNA gene chromatin from S. cerevisiae, providing a detailed description of chromatin at different functional ribosomal DNA (rDNA) elements. Results obtained with the biochemically purified rDNA chromatin were verified by methods analyzing rDNA chromatin composition in vivo. In the next funding period we will use the combination of these approaches to obtain a detailed molecular description of dynamic rDNA chromatin transitions in different physiological situations. We expect to deepen our insights in the structure-function relationship between chromatin and transcription.
Click her for more information.
Project A6: K. Grasser
Transcript elongation and RNA processing/export in Arabidopsis
Our focus will be on elucidating the mechanism(s) of plant nucleocytosolic mRNA transport and identifying molecular links between transcriptional elongation and mRNA export. We plan investigating (1) how export factors are recruited co-transcriptionally to the mRNP and (2) the functional consequences of the association of different export factors with the RNA helicase UAP56. As UAP56 associated factors ALY/UIEF are diversified in plants, (3) we study their functional specificity/redundancy. (4) We intend to identify the plant mRNA export receptor and elucidate its role in the formation of export-competent mRNPs.
Click here for more information.
Project A7: D. Grohmann
Dynamics of transcriptional complexes and interplay of the archaeal RNA polymerase with RNA processing proteins
We plan to investigate the elongation phase of archaeal and eukaryotic transcription. We aim to i) delineate the structural (re-)organisation of the RNA polymerase I transcription machinery during the transition from the initiation to elongation phase at the single-molecule level and ii) to elucidate how transcription and co-transcriptional processes are orchestrated in Archaea with a special focus on the biochemical and structural characterisation of the coupling transcription and translation machinery and the role of Lsm-containing RNPs.
Click here for more information.
Project A8: C. Engel
Structural basis of RNA polymerase I transcription elongation
RNA polymerase (Pol) I consists of 14 protein subunits with a total molecular weight of almost 600 kDa in yeast. Transcription activation by Pol I is accompanied by a contraction of a DNA-binding active center cleft. Here, we will investigate whether activation by contraction and de-activation by expansion are conserved features of Pol I. We will furthermore determine the influence of Pol-I-specific subunits and their domains on the process. To achieve this goal, we employ a single-particle cryo-Electron Microscopy approach.
Click here for more information.
Project A2 (completed in first funding period): P. Cramer
Structure and function of RNA polymerase I and initiation factor Rrn3
The first step in eukaryotic ribosome biogenesis is the production of the ribosomal RNA precursor by RNA polymerase I (Pol I). Pol I is a 14-subunit multi-protein complex with a molecular weight of 600 kDa. The regulation of Pol I and ribosome biogenesis controls cell growth. Despite its importance, the structure of Pol I and the mechanism of class I promoter-specific transcription initiation remain unknown. The long-term goal of this project is to provide the structural basis of Pol I transcription, and Pol I-specific transcription initiation, to understand key switches that control cell growth. In the past, we have determined crystal structures of Pol I subcomplexes A14/43 and A43/34.5 (Geiger et al., Mol. Cell 2010) and the hybrid structure of Pol I that used cryo-electron microscopy, modelling, and biochemical probing, to unravel the functional architecture of the complex (Kuhn, Cell 2007). During the first funding period of this SFB, we propose to determine the X-ray crystal structure of the complete yeast Pol I (Aim 1), and to determine the structure and function of the Pol I-specific initiation factor Rrn3, including its interaction with Pol I (Aim 2). In collaboration with other teams in the SFB, we will also improve crystals of the initiation complex of the archaeal Pyrococcus furiosus polymerase, provide assistance to teams who aim to crystallize factors involved in ribosome biogenesis, and provide help with structure-based prediction and interpretation of site-directed mutagenesis.
Project A3 (completed in first funding period): M. Thomm
Structure function relationships in the archaeal transcription machinery
The archaeal transcription machinery is the evolutionary precursor and an excellent simplified model of the more complex eukaryotic transcription systems. The reconstituted RNA polymerase (RNAP) from Pyrococcus furiosus was a useful tool for the elucidation of major functions of key loops in the active centre of RNAP and the archaeal system was also helpful to unravel a function of the linker region of TFB and of the clamp coiled coil domain of RNAP in open complex formation. Eukaryotic RNAP subunits were shown to functionally replace subunits in the archaeal RNAP, and the archaeal subunit P in WT sequence and a mutated form of subunit H were able to complement deletion mutants of the general eukaryotic RNAP subunits Rpb12 and of the C-terminal domain of subunit Rpb5 in yeast. In this project we aim to analyze the architecture and dynamics of the preinitiation complex (PIC) in more detail in particular with respect to the location and function of different functional elements of TFB that are inserted in the RNA polymerase cleft in a TFIIB-RNA polymerase II cocrystal. We are specifically interested in the transitions from i) closed to open complexes and from ii) open to early elongating complexes. Furthermore, we continue to determine the location of TFE on the RNA-polymerase during initiation and elongation, both in vivo and in vitro. Another target of investigation is transcriptional proofreading of the archaeal RNA polymerase. Reconstitution of the archaeal RNA polymerases allows us to study deletion and point mutants in the trigger loop (TL) to define its contribution to internal cleavage activity of the RNA polymerase, transcriptional fidelity as well as a possible role of TL in interacting with elongation factor TFS, which is located in cocrystals of Pol II and TFIIS close to the TL. These studies will contribute to a deeper understanding of the mechanism of transcription and of the evolution of the three eukaryotic RNA polymerases from a single archaea-like precursor.
Part B: Assembly, function and regulation of RNPs
Project B1: P. Milkereit, H. Tschochner, J. Griesenbeck
Analyses of eukaryotic ribosome assembly
From previous studies we deduce the existence of a number of defined ribosomal protein assembly checkpoints during ribosome maturation in S. cerevisiae. At these checkpoints local assembly states are somehow sensed by specific factors which only promote downstream maturation events if ribosomal protein assembly states in the relevant pre-ribosomal regions were detected as sufficient. We propose to further explore such mechanisms of quality control during ribosome assembly in S. cerevisiae by a combination of structural analyses of ex vivo purified misassembled ribosomal precursors and of structure guided generation of factor and ribosomal protein variants with downstream phenotype analyses.
Click here for more information.
Project B3/B4: G. Meister
Post-translational regulation of small RNA-guided gene silencing pathways
MicroRNA (miRNA)-guided gene silencing is a central process in almost all cellular pathways and therefore affected in many diseases. MiRNAs guide Argonaute (Ago) and TNRC6 proteins to complementary sites on target mRNA leading to gene silencing. This process is embedded into extended regulatory networks including protein interactors as well as post-translational modification pathways. In project B3, we will investigate signaling pathways that affect gene silencing and functionally characterize protein interaction partners of Ago and TNRC6, which we have identified during the pervious funding period. We have already established biochemical purification strategies to isolate Ago- and TNRC6-containg mRNA complex. This will be further optimized and used for structural studies using cryo-electron-microscopy.
Click here for more information.
Project B5: S. Sprunck
Gametogenesis-related small non-coding RNAs and Argonaute proteins in Arabidopsis
Small RNAs and Argonautes are central to gene regulation but analysis of these ribonucleoprotein (RNP) complexes in the flowering plant egg cell remains challenging because it is deeply buried inside the ovary. Project B5 aims to uncover small RNA profiles and RNP complexes of Arabidopsis egg cells and to investigate whether and how miRNAs in the egg cell impact cell specification, fertilization or embryogenesis. Furthermore, project B5 aims to elucidate the molecular mechanism(s) of cell-to-cell small RNA transfer in the microenvironment of the female gametophyte for intercellular communication.
Click here for more information.
Project B6: T. Dresselhaus
Assembly of localized mRNPs and their function in regulating translation in Arabidopsis
The group of Thomas Dresselhaus (B6) aims to study the spatial and temporal control of protein biosynthesis via cytoplasmic mRNPs during early embryo development in Arabidopsis and maize. The multi-step processes of initial mRNP assembly, transport and translational control will be investigated using WOX/WUS mRNA containing mRNPs in early embryos and stem cell niches. They will further study the role of maternal and zygotic RNA-binding proteins and the regulation of their RNPs by DSULylation.
Click here for more information.
Project B9: M.Kretz
Long noncoding RNAs in tissue homeostasis and disease
The project proposed here is designed to analyze the functional impact and modes of action of long non-coding RNAs (lncRNAs) and associated proteins on regulation of epidermal homeostasis in normal and neoplastic tissue. Objective 1: Characterization of the interaction between lncRNA LINC00941 and epigenetic regulator complexes. Objective 2: Analysis of modes of action of LINC00941, SPRR5 and associated proteins in regulation of epidermal differentiation gene clusters. Objective 3: Analyzing functional roles and modes of action of LINC00941, SPRR5 and associated proteins in human tissue neoplasia.
Click here for more information.
Project B10: W. Seufert
Mechanism and regulation of the eIF2-assembly factor Cdc123 and the link of cell cycle entry to mRNA translation
To define the mechanism by which the ATP-grasp protein Cdc123 assembles the hetero-trimeric eIF2 complex we will continue the genetic and biochemical characterization of yeast Cdc123 mutants, employ advanced NMR methods to study protein associations and dynamics, search for a covalent eIF2 modification by SILAC mass spectrometry, and study the in vivo association of eIF2 with the initiator tRNA by RNA immunoprecipitation. To understand the anti-proliferative effect associated with chronic dysfunction of Cdc123 and eIF2, we will profile gene expression, specifically mRNA translation, by RNA-sequencing based methods.
Click here for more information.
Project B11: J. Medenbach
Regulation of alternative translation initiation
The Drosophila RNA-binding protein Sex-lethal (Sxl) is the master regulator of female development in somatic tissues. Numerous regulatory mechanisms operate to tightly control its sex- specific expression and to restrict its activity to female animals. Our aim is to understand in more detail how Sxl expression is regulated and how Sxl and related proteins exert their post-transcriptional regulatory activity on a molecular level.
Click here for more information.
Project B12: R. Sprangers
How do the decapping enzymes DcpS and Dcp2 sense the length of the mRNA substrate?
During mRNA degradation the Dcp2 and DcpS decapping enzymes remove the 5’ mRNA cap structure from the transcript. The substrate for both enzymes is capped RNA, however, Dcp2 is most active on long transcripts, whereas DcpS is only active on short transcripts. Here, we will combine accurate mRNA decapping assays with several biophysical methods to unravel the structural basis for the “molecular rulers” within the Dcp2 and DcpS enzymes. In addition, we will address how the substrate specificity of these enzymes is modulated by interactions with decapping factors and by liquid-liquid phase separations that result in the formation of cellular processing bodies. Our insights have important implications, as the erroneous decapping of an mRNA substrate would lead to significant deregulation of gene expression.
Click here for more information.
Project B13/AP1: S. Ferreira-Cerca
Defining key principles of late small ribosomal subunit assembly in archaea
This project proposal aims to functionally characterise the possible interplay of archaeal specific and conserved bacterial- and eukaryote-like features required for the latest steps of small ribosomal subunit synthesis in archaea. To do so, we aim to functionally characterise selected putative key trans- and cis-acting factors required for the last steps of small ribosomal subunit formation.
Accordingly, this project proposal contributes to the definition of key principles of the in vivo ribosome assembly pathway in archaea and shed light on the intricate evolutionary relationship of the ribosome assembly processes.
Click here for more information.
Project B14: T. Heise & G. Sommer
Biogenesis and Function of RNA Polymerase III transcript-derived piRNAs in Neuroblastoma
RNA-induced gene silencing of transposable elements by piRNA (PIWI-interacting RNA) protects the genomic integrity of germ cells. More recently, the expression of piRNA and PIWI proteins has been associated with cancer, however, the mechanism and function of piRNA biogenesis and piRNA-induced gene silencing in cancer is not well understood. Our preliminary studies demonstrate that cancer-associated RNA-binding protein La (LARP3) is overexpressed in neuroblastoma and that shRNA-mediated depletion of La, which binds to the 3’-end of all RNA Polymerase (Pol) III transcripts, increases the expression of Pol III transcript-derived piRNAs in neuroblastoma cells. Furthermore, RNA-Seq revealed that the increase of those piRNAs correlates with reduced expression of predicted target mRNAs. In our model overexpression of La protein diminishes RNA Pol III transcript-derived piRNA biogenesis, reduces piRNA-induced gene silencing and consequently promotes cancer progression. Hence, this new project aims to study the biogenesis of piRNA, the piRNP composition, and the cellular impact of piRNA RNPs on gene expression in neuroblastoma cells.
Project B7 (completed in first funding period): A. Bosserhoff
Functional implication of miRNA processing in malignant melanoma
Regulation of gene expression by miRNAs plays an important and still emerging role in physiological as well as pathophysiological processes. Details of miRNA function including the characterization of miRNPs still need to be examined to yield a complete understanding of the molecular processes. We have chosen to analyze changes in the miRNA processing machinery in melanoma. These will be determined in detail and effects of these variances will be studied. Preliminary data hinted to deregulation of Argonaute 2 (Ago2) expression in melanoma tumor cells. Therefore, regulation of Ago2 and effects of regulated Ago2 expression will be analyzed in detail. Additionally, we will concentrate on the miRNP and define the influence of protein methylation on miRNP formation and activity. Here, melanoma is an appealing model system as accumulation of a protein-methyltransferase inhibitor, MTA, was shown during cancer development. In summary, successful accomplishment of the aims will yield in a better understanding of the pathophysiological situation of miRNA processing in human disease but also to a broader knowledge on physiological processes.
Z - Service Projects
Project Z1: A. Bruckmann
Mass spectrometry of proteins
In collaboration with different CRC projects we will apply quantitative proteomics methods such as SILAC, iTRAQ, Label-free, and further optimize workflows for our Bruker MaXis plus UHR-QTOF platform. Moreover, we will apply Selected Reaction Monitoring (SRM) as a very sensitive targeted proteomics approach for relative and absolute quantification of proteins and posttranslational modifications. We furthermore aim to implement emerging mass spectrometric methods such as Data-independent acquisition (DIA) for quantitative proteomics and Top-down proteomics for proteoform profiling.
Project Z2: G. Längst
Bioinformatics and advanced RNA/DNA-Seq
The experimental assessment of genome wide experiments requires a sophisticated infrastructure in sequencing equipment and an established platform for the basic analysis of large high throughput datasets. For this we have now established a sequencing and computational core facility in Regensburg, matching the basic needs of the SFB projects. Furthermore, this project is essential in advising and planning genome wide projects within the SFB and requires experienced bioinformaticians to adapt project specific experimental setups and to develop and test novel software pipelines, accordingly.
Project Z3: H. Tschochner
Central Tasks
Graduate Research Academy "RNA Biology"
Graduate Research Academy "RNA Biology"
The scientific personnel hired by the CRC consist mainly of PhD students (currently 23 students). PhD students at the Faculty of Biology and Preclinical Medicine of Regensburg University have to subscribe to the Regensburg International Graduate School of Life Sciences (RIGeL: www.rigel-regensburg.de), which requests students to pursue a structured PhD program including the attendance of summer schools, lab courses, journal clubs, conferences etc., and contains quality control instruments to regulate and monitor the progress of the individual PhD work. RIGeL is subdivided into four disciplines: (i) Cellular Biochemistry and Biophysics, (ii) Molecular Ecology and Evolution, (iii) Neurobiology and (iv) Biomedicine. PhD students of the CRC subscribe to the RIGeL section Cellular Biochemistry and Biophysics (CBB).
RIGeL was established 2009. At the beginning, it has not offered a specific program. Therefore, during the first funding period of the CRC, we have established the CRC-integrated Graduate Research Academy RNA Biology (GRA: www.rnabiology-regensburg.de), which offered a very attractive PhD program besides the RIGeL curriculum to generate the next generation of highly motivated and capable young researchers in the field of RNA biology/biochemistry. In the meantime, the GRA program was extended and improved, and now represents a branch of excellence within the CBB section of RIGeL attracting very good PhD student candidates. Currently, PhD Research Interest Groups in Bioinformatics and cryoEM are formed, which shall actively contribute to the development and problem solving of most recent CRC methodology.
At present 28 PhD students are subscribed to the GRA consisting to 2/3 of CRC PhD students and to 1/3 of PhD students working on related topics, who are not funded by the CRC. In the next funding period we aim to continue the very successful program and offer method as well as generic skills courses, organize annual summer academies, invite alumni speakers and speakers selected by the PhD students, and support PhD students of the various Interest Groups. The next generation of PhD students interested in RNA biology will be motivated and supported to spend, for example, up to three months in a collaborating lab. The GRA also offers recruitment support to PIs to invite, evaluate and eventually hire talented students from foreign countries. In order to attract the most ambitious and outstanding graduate students, a highly attractive structured PhD program is nowadays requested to prepare students both for a non-academic and especially academic career. The CRC thus will strongly benefit not only from the recruitment of talents, but also from students that are given possibilities to attend international method courses and spend short time periods in other labs, thus providing PhD students to initiate networking and contacts with leaders in the RNA world. Moreover, PhD students are also provided with the possibility to propose and hire their own student helpers (WHK) supporting them during their PhD work. In summary the GRA strongly supports highly capable PhD students to become well-educated, critical thinking, problem solving, independent working, responsible and successful individuals. It is expected that PhD students take over responsibilities, shaping and creating the GRA and the CRC in general.
Associated Projects
AP2: J. Perez-Fernandez
Structural and functional analysis of early acting ribosome biogenesis factors
Ribosomes are large cell structures formed by two subunits composed of proteins and RNAs. Cells require production of ribosomes to synthesize proteins and for this reason ribosomes are inherent to cell live. Apparently, the malfunction of ribosomes is the etiology of a set of diseases grouped as ribosomopathies. Thus, it is important to maintain a balanced production of both ribosomal subunits. Production of ribosomes needs to be tightly regulated since ribosomes are dominant constituents of cells and their production consumes a significant amount of cell resources. Since active dividing cells need to duplicate their ribosomal content during the cell cycle, the inhibition of ribosomal synthesis is one of the key processes to counteract cell proliferation in tumor cells.
During the early events of ribosome biogenesis in Saccharomyces cerevisiae, the hierarchical recruitment of more than 40 AFs results in formation of the small ribosomal subunit (SSU) processome. With most of the SSU processome components identified, the current challenges are to understand the mechanisms coordinating transcription of pre-rRNA with ribosome assembly and the mechanisms used by AFs to promote folding and processing of the rRNA. Several AFs associate to form the protein complexes t-UTP/UTP-A, UTP-B, and U3 snoRNP that are primary binders of the pre-rRNA. Still unknown is how these protein complexes nucleate the association of other AFs and initially bundle distant regions of the pre-rRNA. Since formation of the SSU-processome occurs co-transcriptionally, we want to explore possible connections between the polymerase machinery and AFs or pre-rRNA structures that might coordinate transcription and assembly. It would be striking that changes in the elongation rate of polymerase might affect the correct assembly of ribosomal particles by impairing the timing between appearance of RNA sequences and their association with AFs. To solve this conundrum, we propose to study: i) Molecular role of the t-UTP complex during ribosome biogenesis, ii) Role of Pol I transcription during assembly of the SSU-processome and pre-rRNA processing.
Since production of ribosomes constitutes a paradigm to unveil the principles for the biogenesis of ribonucleoprotein complexes (RNP), the biochemical principles behind ribosome assembly could be of general interest for biogenesis of other RNPs.
Click here for more information.
AP 3: A. Bleckmann
Selective mRNA localisation at the endoplasmic reticulum
The endoplasmic reticulum (ER) is a large, continuous membrane-bound organelle comprised of functionally and structurally distinct domains. Three major morphology domains can be distinguished as (i) nuclear envelope, (ii) peripheral ER cisternae, and (iii) an interconnected tubular network. Proteins which are destined for secretion or localize to the secretion pathway are translated at ER structures decorated with Ribosomes and are thus called the rough ER. Ribosomes are trafficking to the ER via the signal-recognition particle pathway, when synthesising proteins that contain a specific signal sequence. Than the proteins are distributed by the secretory pathway to their destination. If proteins need to be localised in a polar manner, the current model proposes that those proteins are distributed polar by the secretory pathway, while localisation of translation is unaffected.
However, synergid cells, which are pollen tubes attracting cells of the female gametophyte, secrete molecules at one pole of the cell in order to create a signal gradient, guiding the pollen tube into the female gametophyte. This makes synergid cells a great model to study polar translation and secretion mechanisms. On the level of mRNA translation and localisation, we could observe a polar distribution of mRNAs towards the filiform apparatus. However, the ER does not show this accumulation at one site of the cell. Therefore, we propose that the polar localisation of a translated mRNA might be due to compartmentalization of the ER. This mechanism is likely induced by an unknown sporophytic trigger, as we see mRNA polarization only after pistil pollination, but before the male pollen tubes reach the synergid cells itself.
With support of the SFP960 and various collaborations we would like to analyse the ribosome distribution at the ER in response to the fertilisation process and identify the trigger responsible for those global changes. Notably, a special feature of plants is a high heterogeneity in ribosomal populations. In Arabidopsis, each of the approximately 80 ribosomal proteins is encoded by two to seven paralogs, which are all in-cooperated into functional ribosomes, depending on tissue und developmental-specific expression pattern as well as on environmental triggers. Therefore, we are currently searching for synergid-specific ribosomal proteins and analyse their subcellular localisation during the process of fertilisation. Additionally, we are analysing proteins of the signal recognition particle, which are also represent by several paralogues in the Arabidopsis genome. Studying those two RNP complexes will give us some insight into the polar localisation of translated mRNA.
Click here for more information.