Forschung
This research group focuses on:
The integration of light-based technologies to construct ‘smart’ systems that are greater than the sum of their parts.
Crafting robust systems that merge cutting-edge hardware and software to enhance performance, efficiency, and reliability for biomedical imaging and analysis.
Development of imaging and spectroscopy systems in interdisciplinary settings, demonstrating integration innovations across optical physics, mass spectrometry and bioinformatics.
Exploring physically explainable AI for label-free imaging of fast-moving cells.
Development of novel spectral imaging methods (Brillouin, Raman, Ion spectroscopy) for biomedical research, Bioinformatics and ML approaches to multi-modal data integration.
Explainable and physics-aware artificial intelligence
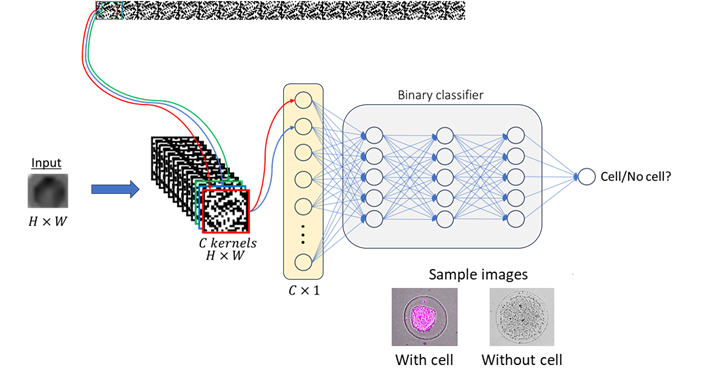
Flow cytometry is a key technology in cell biology, enabling high-throughput analysis of cell populations at up to 100,000 particles per second. Traditional methods rely on fluorescent labels to detect specific proteins, requiring carefully designed staining panels which is still fundamentally limited by spectral overlap. To expand detection capabilities, label-free approaches have been explored, primarily using Raman spectroscopy. However, Raman-based methods lack quantitative molecular data and have significantly lower throughput than labelled techniques.
To address these limitations, we propose a novel high-speed, label-free cell analysis method for flow cytometry. Our approach leverages only single-pixel photodetectors (e.g., photodiodes) and the rapid motion of droplets. As a droplet moves through a region of spatially modulated light, its biophysical properties (e.g., shape, size) are encoded in the photodiode signal. These signals are then processed using deep learning models trained to extract meaningful cellular features. We employ convolutional neural networks (CNNs) to analyse the photodiode signal, learning spatial and temporal patterns that distinguish between cell-containing and empty droplets. By training with a mixture of simulated and experimental data, initial results demonstrate that our deep learning framework can accurately classify the presence of cells in droplets, setting the foundation for high-throughput, label-free multi-omics analysis.
Development of novel spectral imaging methods for biomedical research
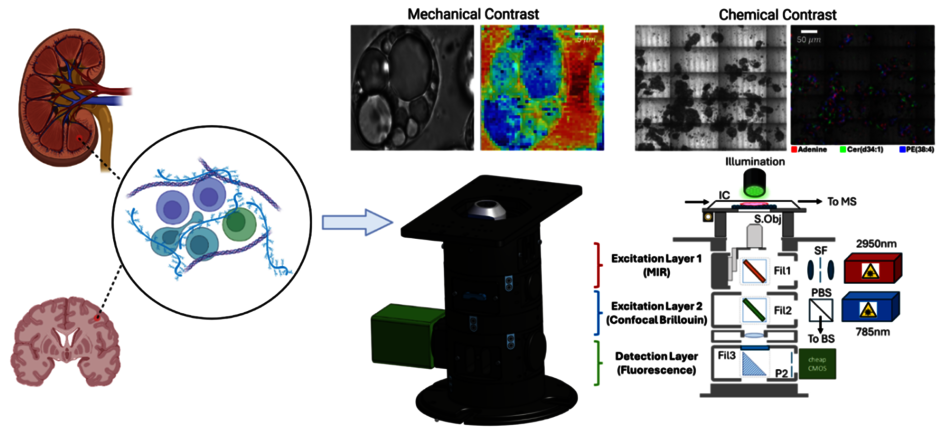
Pathogenesis is the process by which a disease begins and develops in the body, from the initial cause to the symptoms it produces. Although conditions affect different organs, they share many underlying features and mechanisms.
While most research has focused on immune genes and signalling, growing evidence shows that the physical properties of tissues — such as stiffness — strongly influence how immune cells move, activate, and communicate, suggesting that changes in tissue mechanics can trigger and drive disease. We thus propose that mechanobiology is a critical but underexplored driver of autoimmune disease.
To understand how physical forces shape various diseases e.g., cancer, we have developed label-free, spectral imaging techniques like special camera that can detect and map stiffness and chemistry directly in intact tissues, allowing us to reveal how immune cells sense and respond to their physical environment. By imaging these mechanochemical interactions in their native environment, we will identify shared pathways with diagnostic or therapeutic potential and uncover disease-specific mechanical signatures.
Bioinformatics and ML approaches to multi-modal data integration
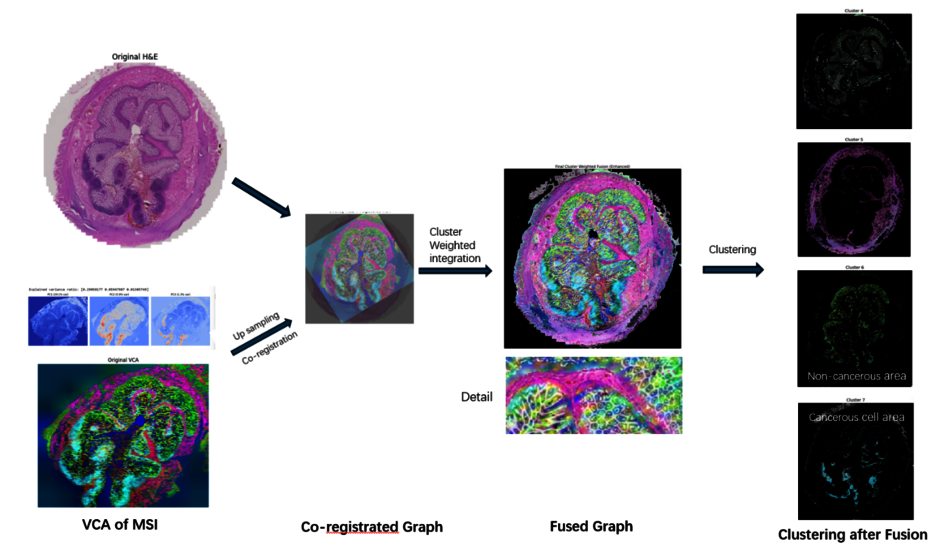
Early detection of many diseases is crucial for improving patient outcomes, but the complex, multifactorial nature of some major remaining challenges to human health e.g., Alzheimer’s, has hindered the discovery of reliable early-stage biomarkers. In simple terms, one modality used to study the disease provides only a partial view of the disease landscape. Integrating complementary spatial modalities—such as magnetic resonance imaging (MRI), positron emission tomography (PET), and immunofluorescence microscopy—enables a more comprehensive understanding of pathology by combining molecular, structural, and functional information. However, multimodal spatial integration presents several analytical challenges.
This project employs and potentially enhances cutting-edge manifold learning techniques, including:
UMAP (Uniform Manifold Approximation and Projection)
Diffusion Maps
Custom graph-based algorithms
These methods will allow us to:
Reduce the dimensionality of complex datasets
Preserve crucial spatial and molecular information
Uncover hidden patterns and relationships in the data
