Cell Cloud Analysis
The presence or absence of specialized immune cells can cause disease or affect treatment response. If the characteristics of these cells are unknown, artificial intelligence can help defining and characterizing these critical cells. We develop an automated analysis pipeline based on convolutional networks already in use in the field of point cloud classification to (a) identify these cells from flow cytometry data and (b) to predict treatment response.This project is carried out in close collaboration with INsTRuCT.
Metabodefense: The host metabolism as an antimicrobial effector
Products of the body’s own metabolism not only have a regulatory effect on the immune system but can also influence the growth or persistence of bacteria. Metabodefense aims at finding antimicrobial structures within the host metabolism that enhance pathogen control. To this end, we apply machine learning techniques to predict pathogen control and causal discovery algorithms to identify potential molecular targets. Furthermore, we work on preprocessing methods for high-throughput metabolic data sets based on Mixed Graphical Models which can be used to detect anomalous data points in high dimensional data. This project is part of bayresq.net (externer Link, öffnet neues Fenster)
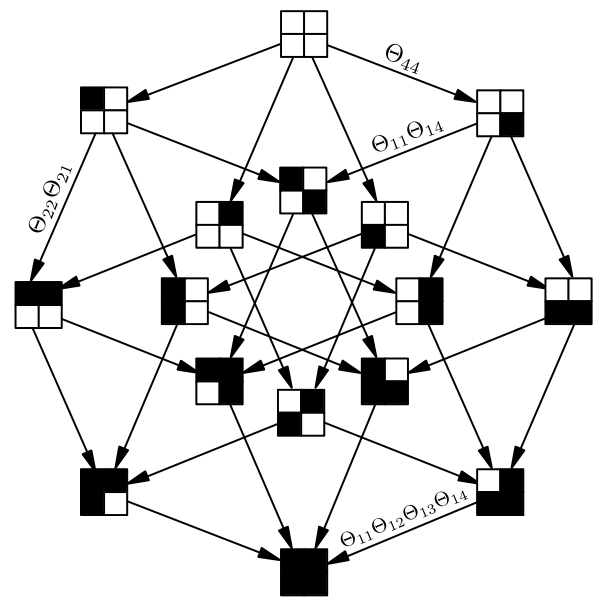
Tensor Approximations in Computational Oncology
Tumors progress through the accumulation of mutations in their genome and this process can be modelled as a Markov Chain on the space of all possible genotypes. For n mutations this space has size 2^n , and the rate matrix of the process holds 2^n x 2^n elements. For more than n=130 mutations this matrix holds more elements than there are atoms in the universe. To compute the likelihood of such a model the matrix must be inverted. This is what we aim to do in this project. We use low dimensional parametrizations for the rate matrix that allows to store it in strongly compressed formats such as tensor trains or in the hierarchical Tucker format. In this project we collaborate with the groups of Lars Grasedyck (externer Link, öffnet neues Fenster) and Tilo Wettig (externer Link, öffnet neues Fenster)
AI assisted lymphoma pathology
Oncologic precision medicine needs very exact sub-typing of tumors to enable the best possible therapy response. While pathologist are very good at this, diagnosing these tumors is a very laborious and analog process with limits in how many patients can be diagnosed, how much area of the slides can be looked at and how many subtypes are distinguishable by the human eye. But augmenting this process through digitalisation is possible, slides can be scanned digitally as images with about 10 gigapixels resolution. We are developing a solution to automatically diagnose different subtypes of lymphoma or benign lymphadenitis based on these high resolution scans of lymph node slides. We use deep convolutional neural networks to train a pathology assistance tool based on a data set of 628 scanned and labeled slides from 157 patients with 4 different immunohistochemistry stains each.
Zero-Sum Regression
Molecular measurements are relative to a reference point, like a fixed aliquot of RNA extracted from a tissue, a defined number of blood cells, or a defined volume of biofluid. Reference point discrepancies across data sets compromise the performance of regression models like the LASSO. We have developed zero-sum regression for a reference point insensitive analysis that enhance cross-platform analysis. An R-package implementation can be downloaded here (externer Link, öffnet neues Fenster)
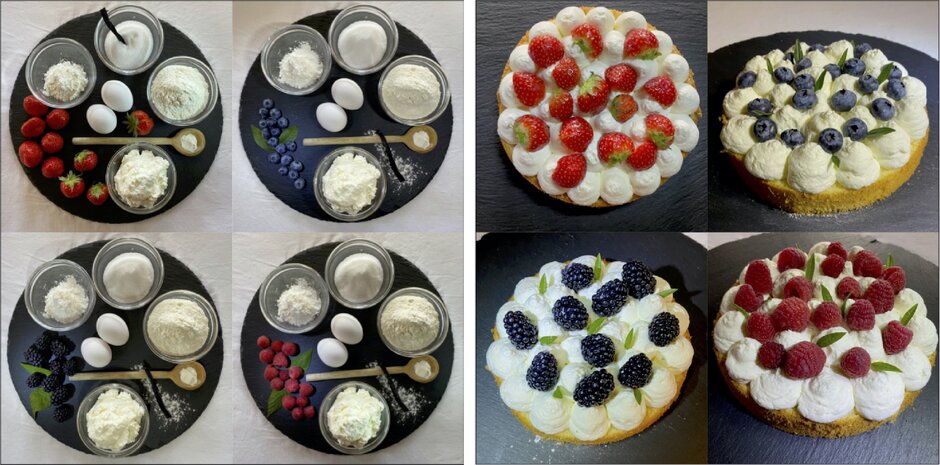
Digital Tissue Deconvolution
A bulk tumor tissue consists of a heterogeneity of multiple cells. As malignant and immune cells are embedded into the microenvironment, a bulk is a merger of multiple elements - just as a cake. Deconvoluting a tumor tissue leads to its cellular composition, deconvoluting a cake leads to its ingredients. As it is not that easy to taste every ingridient and its amount, it is not that easy to estimate the cellular composition of tumor gene expression measurement.
For that task (deconvolution a tumour bulk, not the cake), we developed the algorithm Digital Tissue Deconvolution (DTD), which predicts the cellular composition given a expression measurement. DTD increaes deconvolution accuracy by adapting a deconvolution model to the tissue scenario. Check out our github page (externer Link, öffnet neues Fenster) , or our publications Schoen et al. 2020 (externer Link, öffnet neues Fenster) and Goertler et al. 2020 (externer Link, öffnet neues Fenster)

TissueResolver - a new algorithm to predict cell type specific gene expression in tissues
From the many available single cell profiles publicly available, we find and combine a small number of "prototypic" cells that are best suited to describe RNASeq data of bulk tissue, allowing us to attribute gene expression to certain sub-populations of cells and address the question "what are the individual cells in the tissue doing?". This approach preserves the biological complexity of cells by avoiding model assumptions but turns the problem into a combinatorial machine learning problem that is NP-hard and requires smart and efficient algorithms."
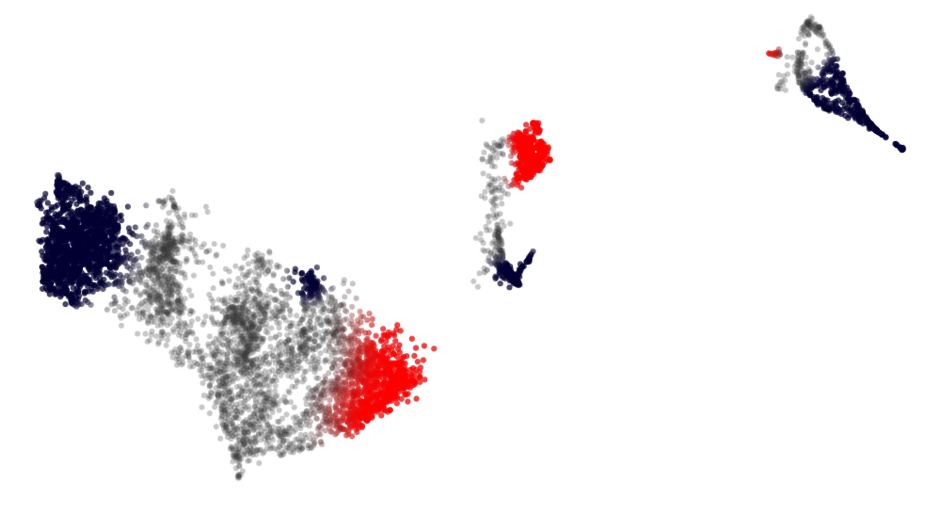

Celloscope
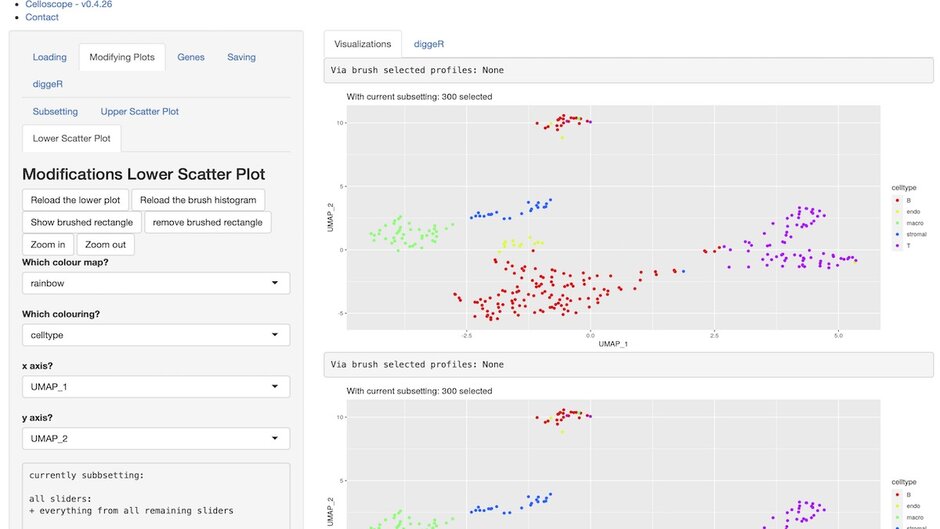
Preprocessing a scRNA-Seq dataset – including calculating various quality metrics, finding suitable parameters for 2D embeddings via e.g., UMAP and clustering for cell type calling – takes time. Changing parameters, subsetting for another QC metric, and always visually observing whether the biological connection between the profiles is still valid, drains time.
Quality assessing, including selecting thresholds for QC metrics, of scRNA-Seq profiles does not require automated scripts, but an easy-to-use application – as it must be redone on each and every dataset potentially multiple times.
We present celloscope , a shiny-based web application that easily handles preprocessing and visualization of scRNA-Seq datasets. Celloscope visualizes all potential information of a data set directly in the browser. It solves as a clickable and intuitive alternative to automated scripts, without losing functionality.
Visit celloscope.spang-lab.de (externer Link, öffnet neues Fenster) , or checkout our github page (externer Link, öffnet neues Fenster)